Overview
Protein-sequence similarity search is at the centre of a lot of modern biotech work — drug-target discovery, function prediction, evolutionary analysis. The standard tool for the job is BLAST, and BLAST is accurate, but on a database with tens of millions of sequences a single query can take minutes to hours. That latency makes interactive exploration impossible and forces researchers to batch queries overnight.
The biotech R&D team that brought us in wanted what everyone wants: BLAST-grade recall at interactive speed. Researchers should be able to type in a sequence, get neighbours back in under a second, and click through them to explore the surrounding evolutionary or functional landscape without ever staring at a progress bar.
ForthClover designed and shipped a domain-aware embedding-based similarity-search system on AWS. It indexes more than 10M protein sequences in a single pane and returns the top neighbours for any query in less than 100 ms — roughly three orders of magnitude faster than the team's previous BLAST-based pipeline at comparable recall.
AWS architecture
The system has three responsibilities: turn raw protein sequences into useful embeddings, store and retrieve those embeddings with sub-second latency, and serve search results to research clients through a clean API.
Embedding & inference layer
- Protein-domain embedding model: a model tuned specifically for biological sequence semantics rather than generic text similarity, so "close" in vector space genuinely correlates with biological closeness.
- Amazon EC2 GPU instances: on-demand and spot-mixed GPU fleet handling embedding generation for both the corpus build and live query embedding.
- Batch + streaming pipelines: the corpus is embedded in batch; new sequences are embedded continuously as they arrive in the team's upstream data store.
Indexing & retrieval layer
- High-performance vector index: hybrid inverted-file + HNSW configuration sized for the working set, deployed on memory-optimised instances.
- Hybrid search: vector similarity is combined with structural filters (taxonomy, length range, functional annotation) so researchers can narrow before they reach the embedding step.
- Caching layer: Redis-backed cache for the most-queried sequences and pre-computed neighbour sets, taking the long tail of repeat lookups off the index.
API & delivery layer
- RESTful API: single, narrow query surface for client tools, designed so existing notebooks and internal apps can integrate in a few lines of code.
- Auto-scaling: API tier scales with query load so quiet hours stay cheap and big research pushes don't queue.
- Observability: per-query latency, recall, and index-health metrics surfaced to a Grafana dashboard the platform team owns.
Benefits
The benefits split cleanly across three groups: researchers, the platform team that runs the system, and the science itself.
Performance
Traditional vs embedding-based search
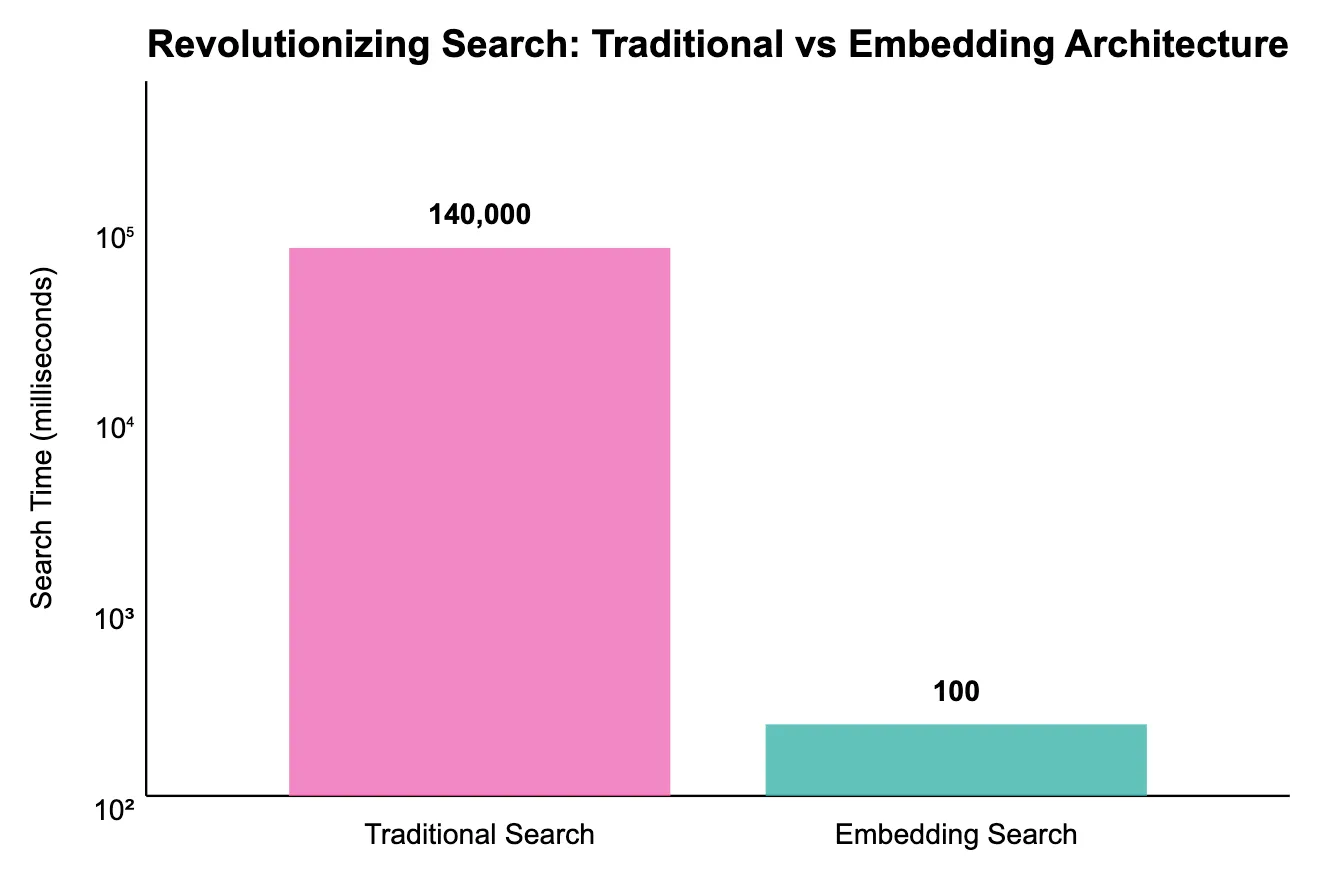
Median query latency on the same 10M-sequence corpus — embedding-based search returns neighbours in under 100 ms versus minutes-to-hours for the BLAST baseline.
Performance
- < 100 ms median search: across a corpus of more than 10 million protein sequences.
- ~1000× faster than BLAST at comparable recall on the team's benchmark queries.
- Interactive exploration: researchers can click through neighbour sets in real time instead of queueing batch jobs.
- Stable under load: p95 latency stays within budget even during periods of concurrent research activity.
Operational characteristics
- Predictable cost: mix of reserved and spot GPU capacity for embeddings, memory-optimised on-demand for the index. No standing GPU cluster.
- Cleanly scaled: the index can grow into the next 10M sequences without re-architecting.
- Reproducible: the entire stack lives in Terraform, so spinning up a staging copy is a single command.
- Owned by the platform team: no novel services for them to learn — every component is either AWS or a familiar open-source primitive.
Research impact
- Drug-target discovery: researchers can sweep through analogues for a known target in minutes instead of days.
- Function prediction: unknown sequences are placed against known-function neighbours interactively.
- Evolutionary analysis: cross-species relationships surface as part of routine querying rather than as a separate study.
- Structure-aware lookups: hybrid filters let researchers combine functional and structural signals in a single query.
What changes for a research team when similarity search drops from minutes to milliseconds is not just speed — it is a different way of working. The BLAST-era workflow was batch-and-wait. The new workflow is type-and-explore. ForthClover was glad to partner with the team to design an AWS-native architecture that makes that interactive workflow possible at the scale their corpus actually requires.